UT Southwestern researchers have identified vast webs of small snippets of the genome that interact with each other and with genes to promote prostate cancer. Their findings, published June 22 in the Journal of Clinical Investigation, could lead to new ways to treat the most common type of malignancy in American men other than skin cancer.
Research over the past few decades has shown that genetic mutations trigger and encourage the growth and spread of many cancer types. However, prostate cancers typically have relatively few of these mutations, explains study leader Ram S. Mani, Ph.D., an assistant professor in the departments of pathology and urology at UTSW.
Instead, how prostate cancers regulate gene expression—which genes are turned off and on, and to what extent—tends to go awry. What causes this aberrant gene regulation has been a mystery. Although previous research had identified pieces of DNA throughout the genome that appeared to be connected to prostate cancer, many of these pieces weren’t genes, so their role was unclear.
To help answer these questions, Mani and his colleagues focused on genetic enhancers, short pieces of DNA that help encourage genes to make proteins. However, explains Mani, it’s not as simple as a one-to-one ratio of enhancer per gene target. One enhancer may target several genes, or multiple enhancers might target a single gene. Additionally, each enhancer could be up to a million base pairs away from its gene target on the same chromosome—a significant distance that makes it difficult to link enhancers with their genes.
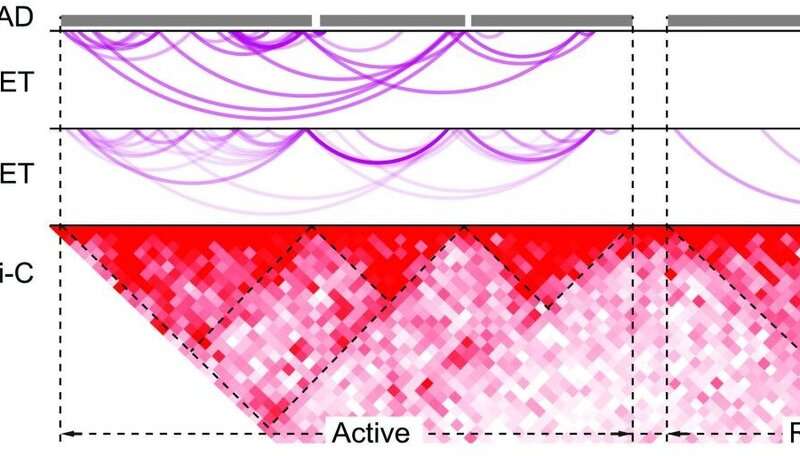
To pair enhancers to their gene targets in prostate cancer, Mani and his team used a technique called ChIA-PET, short for chromatin interaction analysis by paired-end tag sequencing. This technique maps associations of particular proteins with DNA throughout the genome.
The researchers used ChIA-PET to find DNA targeted by a protein called RNA polymerase II, looking for differences between benign prostate cells and prostate cancer cells. RNA polymerase II not only binds to genes to start transcription—the process in which cells use the instructions in DNA to make RNA—but also enhancers for these genes. “When this technique identifies a gene, it also identifies all the enhancers that target that gene at the same time,” says Mani, a member of UT Southwestern’s Harold C. Simmons Comprehensive Cancer Center.
Using sophisticated analytical techniques, the scientists sorted out each of these complicated interactions unique to prostate cancer cells, finding vast hubs of enhancers that interact not just with genes but also with each other. They found that many enhancers interacted with tens of other enhancers or genes. These interactions explain the abnormal expression of several key prostate cancer genes, such as the androgen receptor gene that’s over-expressed in many subtypes of prostate cancer and the oncogene MYC, which plays a role in many cancer types. Further investigation identified some “grammar rules” for enhancers in prostate cancer, grouping genes and enhancers into clusters that strictly interact with each other.
Mani notes that finding these hubs of enhancers and their genes also helps explain the long-standing mystery of the short pieces of DNA connected to prostate cancer that are scattered throughout the genome. This new research suggests that many of these pieces are enhancers.
Source: Read Full Article
