The novel coronavirus pandemic has now resulted in more than 3 million confirmed cases globally and is pushing scientists to share ideas quickly and figure out the best ways to collaborate and contribute to solutions.
Recently, Duke researchers across the School of Medicine came together for an online symposium consisting of several short presentations to summarize the latest of what is known about the novel coronavirus, SARS-CoV-2.
This daylong event was organized by faculty in the Department of Molecular Genetics and Microbiology and researchers from different fields to share what they know about the virus and immunity to guide vaccine design. This conference highlighted the myriad new research pathways that Duke researchers are launching to better understand this pandemic virus.
One neat area of research is understanding viral processes within cells to identify steps at which antivirals may block the virus. Stacy Horner’s Laboratory studies how RNA viruses replicate inside human cells. By figuring out how viruses and cells interact at the molecular level, Horner can inform development of antivirals and strategies to block viral replication. Antivirals stop infections by preventing the virus from generating more of copies of itself and spreading to other cells. This controls damage to our cells and allows the immune system to catch up and clear the infection.
At the symposium, Horner explained how the SARS-CoV viral genome consists of 29,891 ribonucleotides, which are the building blocks of the RNA strand. The viral genome contains 14 areas where the RNA code can be transcribed into shorter RNA sequences for viral protein production. Though each RNA transcript generally contains the code for a single protein, this virus is intriguing in that it uses RNA tricks to code for up to 27 proteins. Horner highlighted two interesting ways that SARS-CoV packs in additional proteins to produce all the necessary components for its replication and assembly into new viral progeny.
The first way is through slippery sequences on the RNA genome of the virus. A ribosome is a machine inside the cell that runs along a string of RNA to translate its code into proteins that have various functions. Each set of 3 ribonucleotides forms one amino acid, a building block of proteins. In turn, a string of amino acids assembles into a distinct structure that gives rise to a functional protein.
One way that SARS-CoV-2 packs in additional proteins is with regions of its RNA genome that make the ribosome machinery slip back by one ribonucleotide. Once the ribosome gets offset it reads a new grouping of 3 ribonucleotides and creates a different amino acid for the same RNA sequence. In this way, SARS-CoV-2 makes multiple proteins from the same piece of RNA and maximizes space on its genome for additional viral proteins.
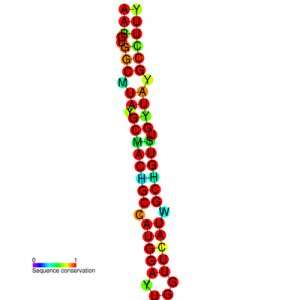
Secondly, the RNA genome of SARS-CoV-2 has regions where the single strand of RNA twists over itself and connects with another segment of RNA farther along the code to form a new protein. These folds create structures that look like diverse trees made of repetitive hairpin-like shapes. If the ribosome runs into a fold, it can hop from one spot in the RNA to another disjoint piece and attach a new string of amino acids instead of the ones directly ahead of it on the linear RNA sequence. This is another way the SARS-CoV-2 packs in extra proteins with the same piece of RNA.
Horner said a step-by-step understanding of what the virus needs to survive at each step of its replication cycle will allow us to design molecules that are able to block these crucial steps.
Indeed, shapes of molecules can determine their function inside the cell. Three Duke teams are pursuing detailed investigation of SARS-CoV-2 protein structures that might guide development of complementarily shaped molecules that can serve as drugs by interfering with viral processes inside cells.
For example the laboratory of Hashim Al-Hashimi, develops computational models to predict the diversity of structures produced by these tree-like RNA folds to identify possible targets for new therapeutics. Currently, the Laboratories of Nicholas Heaton and Claire Smith are teaming up to identify novel restriction factors inside cells that can stop SARS-CoV-2.
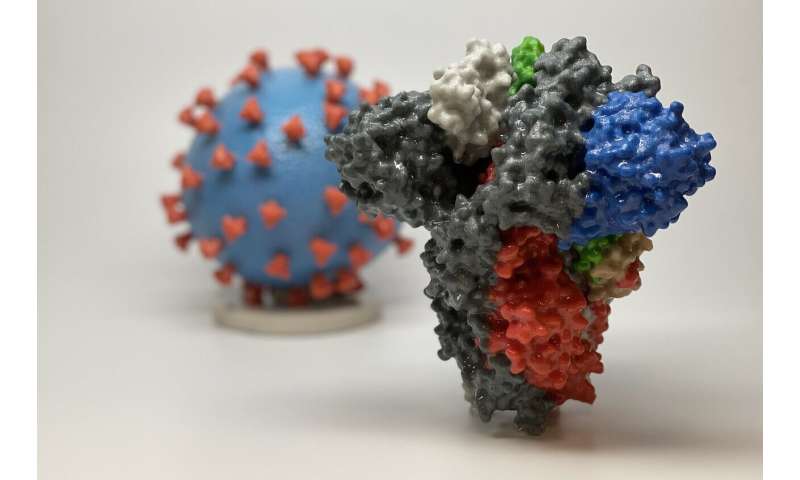
However, it is not just the structures of viral components expressed inside the cells that matter, but also those on the outside of a virus particle. In Latin, corona means a crown or garland, and coronaviruses have been named for their distinctive crown-like spikes that envelop each virus particle. The viral protein that forms this corona is aptly named the “Spike” protein.
This Spike protein on the viral surface connects with a human cell surface protein (Angiotensin-converting enzyme 2, abbreviated as ACE2) to allow the virus to enter our cells and cause an infection. Heaton proposed that molecules designed to block this contact, by blocking either the human cell surface protein or the viral Spike protein, should also be tested as possible therapies.
One promising type of molecule to block this interaction is an antibody. Antibodies are “Y” shaped molecules that are developed as part of the immune response in the body by the second week of coronavirus infection. These molecules can detect viral proteins, bind with them, and prevent viruses from entering cells. Unlike several other components on our immune defense, antibodies are shaped to specifically latch on to one type of virus. Teams of scientists at Duke led by Dr. Sallie Permar, Dr. Georgia Tomaras, and Dr. Genevieve Fouda are working to characterize this antibody response to SARS-CoV-2 infection and identify the types of antibodies that confer protection.
Infectious disease specialist Dr. Chris Woods is leading an effort to test whether plasma with antibodies from people who have recovered can prevent severe coronavirus disease in acutely infected patients.
Source: Read Full Article
