In Huntington’s disease, a faulty protein aggregates in brain cells and eventually kills them. Such protein aggregates could, in principle, be prevented with a heat shock protein. However, it is not well known how these proteins interact with the Huntington’s disease protein. New research by Patrick van der Wel (University of Groningen, the Netherlands) and colleagues at the University of Texas has partially resolved the structure of heat shock proteins that bind to such aggregating proteins, helping us to understand how they work. The results were published on 11 February in the journal Nature Communications.
Heat shock proteins (Hsp) are produced by cells that are exposed to stressful conditions. The Hsp family is diverse, and quite a few of the proteins function as chaperone. This means that they help other proteins to fold (or re-fold after being damaged) in the correct way. “These proteins can assist in folding thousands of different proteins. To this end, they use co-chaperones with specific binding abilities,” explains Patrick van der Wel, Associate Professor of Solid-State NMR Spectroscopy at the University of Groningen.
Ordered
One class of heat shock proteins, Hsp40, helps to suppress protein aggregates like those that appear in Huntington’s disease. These Hsp40 proteins come in different kinds, and some of them will bind specifically to aggregating proteins with a lot of repeated glutamine amino acids, like the faulty protein found in Huntington’s disease. One of these Hsp40 proteins is called DnaJB8, and this was the protein studied by Van der Wel and his colleagues.
“In order to understand the action of DnaJB8, we need to know what it looks like,” says Van der Wel. However, it is difficult to resolve the structure of this type of protein. “It appears as a dimer or an oligomer, so a number of these protein units work together, but their structure is not really ordered,” he continues. This makes it impossible to use standard techniques, which all require ordered structures.
Carbon atoms
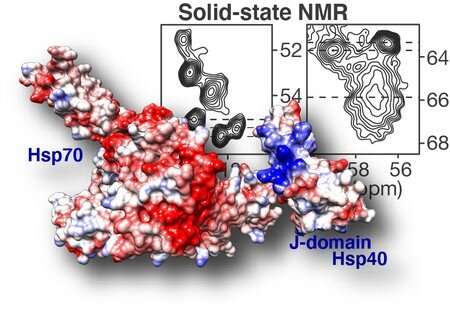
Van der Wel was asked by colleagues at the University of Texas to help tackle this problem. Van der Wel is specialized in solid-state NMR spectroscopy, a technology that can measure how atoms are connected to each other. In simple terms, the NMR signals of two connected carbon atoms in DnaJB8 depend on how they interact with other atoms in the molecule. Therefore, the measured spectrum of the carbon atoms can show in which amino acid they are located. Such information can be used to get an idea of the protein’s structure, even if it is not well ordered.
The DnaJB8 protein is made up of different domains, with different functions. Through a series of experiments, Van der Wel was able to determine which domains are stuck inside the DnaJB8 protein, and which are available on the outside. The experiments suggested that the so-called ‘J domain’ of DnaJB8 was able to switch between being stuck and being accessible. This is important because this part of the DnaJB8 protein is responsible for turning on the Hsp70 protein, which can prevent the protein aggregates from forming. In other words, there seems to be a ‘switch’ in DnaJB8 that controls this interaction with Hsp70. Interestingly, this switch was found to be located in a domain of DnaJB8 whose exact role was previously unclear.
Hypothesis
“Thus, our hypothesis based on the structure was that the DnaJB8 is inactive until it binds to the faulty proteins and that it then attracts Hsp70,” says Van der Wel. A series of simulations and experiments at the University of Texas confirmed this idea and yielded a detailed model of how these proteins work together.
Source: Read Full Article
