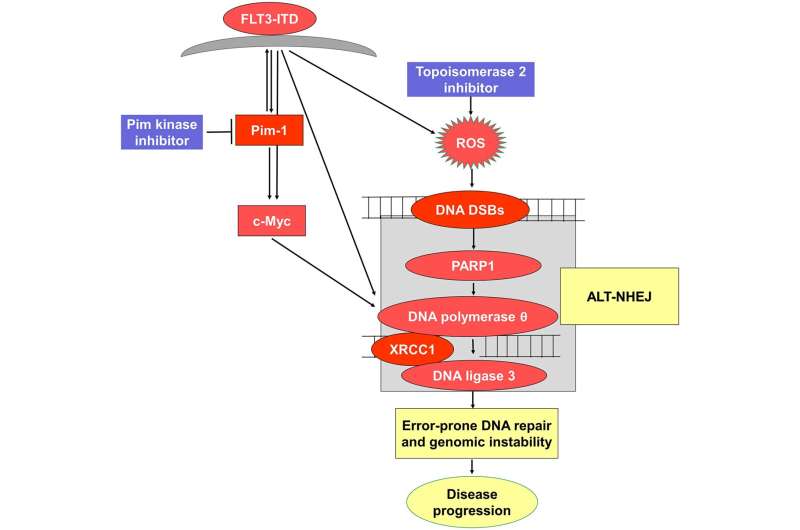
The journal Oncotarget has published “Pim kinase inhibitor co-treatment decreases alternative non-homologous end-joining DNA repair and genomic instability induced by topoisomerase 2 inhibitors in cells with FLT3 internal tandem duplication” which reported that the serine/threonine kinase Pim-1 is upregulated downstream of FLT3-ITD, and inhibiting Pim increases topoisomerase 2 inhibitor chemotherapy drug induction of DNA DSBs and apoptosis. Alt-NHEJ activity, measured with a green fluorescent reporter construct, increased in FLT3-ITD-transfected Ba/F3-ITD cells treated with TOP2 inhibitors, and this increase was abrogated by Pim kinase inhibitor AZD1208 co-treatment.
TOP2 inhibitor and AZD1208 co-treatment downregulated cellular and nuclear expression of c-Myc and Alt-NHEJ repair pathway proteins DNA polymerase, DNA ligase 3 and XRCC1 in FLT3-ITD cell lines and AML patient blasts.
Dr. Maria R. Baer from The University of Maryland as well as The Veterans Affairs Medical Center says that “internal tandem duplication (ITD) of the fms-like tyrosine like kinase 3 (FLT3) receptor tyrosine kinase is present in acute myeloid leukemia (AML) cells of 30 percent of patients, resulting in constitutive and aberrant FLT3 signaling.”
Remission induction chemotherapy including the nucleoside analog cytarabine and an anthracycline and/or other topoisomerase 2 inhibitor remains first-line therapy for AML. While patients with AML with FLT3-ITD generally achieve remission with chemotherapy, they relapse rapidly and AML cells frequently have new structural chromosome abnormalities at relapse, consistent with a role of genomic instability in FLT3-ITD AML progression and relapse.
Inhibition of Pim kinases is therefore an attractive therapeutic approach in AML with FLT3-ITD. Pan-Pim kinase inhibitors are being developed, and several have proceeded to early phase clinical testing; the most recent is TP-3654. HR, mediated by BRCA1, BRCA2 and RAD51 and occurring in the S and G2 phases of the cell cycle, uses the complementary sister chromatid strand as a template for repair and is therefore highly accurate and error-free. C-NHEJ is initiated by the Ku proteins Ku70/80, followed by recruitment of the DNA-dependent protein kinase catalytic subunit, and mediates repair by directly ligating the ends of DSBs, creating the potential for small deletions, insertions and other structural errors at repair sites.
Source: Read Full Article
